Intra-tumoral Microbiome Data Analysis
Trends in Microbiology
Qi Wang, Zhaoqian Liu, Anjun Ma, Zihai Li, Bingqiang Liu, Qin Ma
The human microbiome is intimately related to cancer biology and plays a vital role in the efficacy of cancer treatments, including immunotherapy. Extraordinary evidence has revealed that several microbes influence tumor development through interaction with the host immune system, that is, immuno–oncology–microbiome (IOM). This review focuses on the intratumoral microbiome in IOM and describes the available data and computational methods for discovering biological insights of microbial profiling from host bulk, single-cell, and spatial sequencing data. Critical challenges in data analysis and integration are discussed. Specifically, the microorganisms associated with cancer and cancer treatment in the context of IOM are collected and integrated from the literature. Lastly, we provide our perspectives for future directions in IOM research.
-Qin.png)
bioRxiv
Zhaoqian Liu, Yuhan Sun, Anjun Ma, Xiaoying Wang, Dong Xu, Daniel Spakowics, Qin Ma, Bingqiang Liu
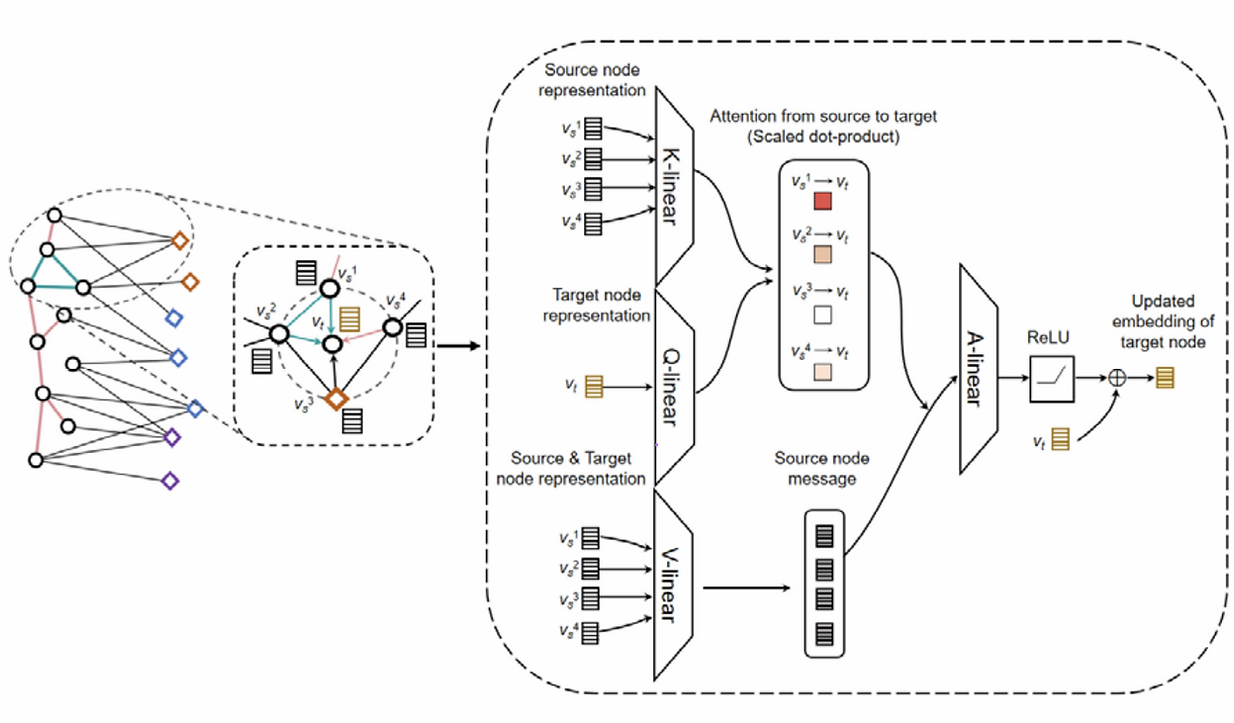
Microbes are extensively present among various cancer tissues and play a vital role in cancer prevention and treatment responses. However, the underlying relationships between intratumoral microbes and tumors are still not well understood. Here, we developed a MIcrobial Cancer-association Analysis using a Heterogeneous graph transformer (MICAH) to identify intratumoral cancer-associated microbial communities. MICAH integrates metabolic and phylogenetic relationships among microbes into a heterogeneous graph representation. It uses a graph attention transformer to holistically capture the relationships between intratumoral microbes and cancer tissues, which improves the explainability of the association between identified microbial communities and cancer. We applied MICAH to intratumoral microbiome data across five cancer types and demonstrated its good generalizability and reproducibility. We believe this graph neural network framework can provide novel insights into cancer pathogenesis associated with the intratumoral microbiome.
Under review
Cankun Wang, Anjun Ma, Megan E. McNutt, Rebecca Hoyd, Caroline E. Wheeler, Lary A. Robinson, Carlos H.F. Chan, Yousef Zakharia, Rebecca D. Dodd, Cornelia M. Ulrich, Sheetal Hardikar, Michelle L. Churchman, Ahmad A. Tarhini, Eric A. Singer, Alexandra P. Ikeguchi, Martin D. McCarter, Nicholas Denko, Gabriel Tinoco, Marium Husain, Ning Jin, Afaf E.G. Osman, Islam Eljilany, Aik Choon Tan, Samuel S. Coleman IV, Louis Denko, Gregory Riedlinger, Bryan P. Schneider, Daniel Spakowicz, Qin Ma
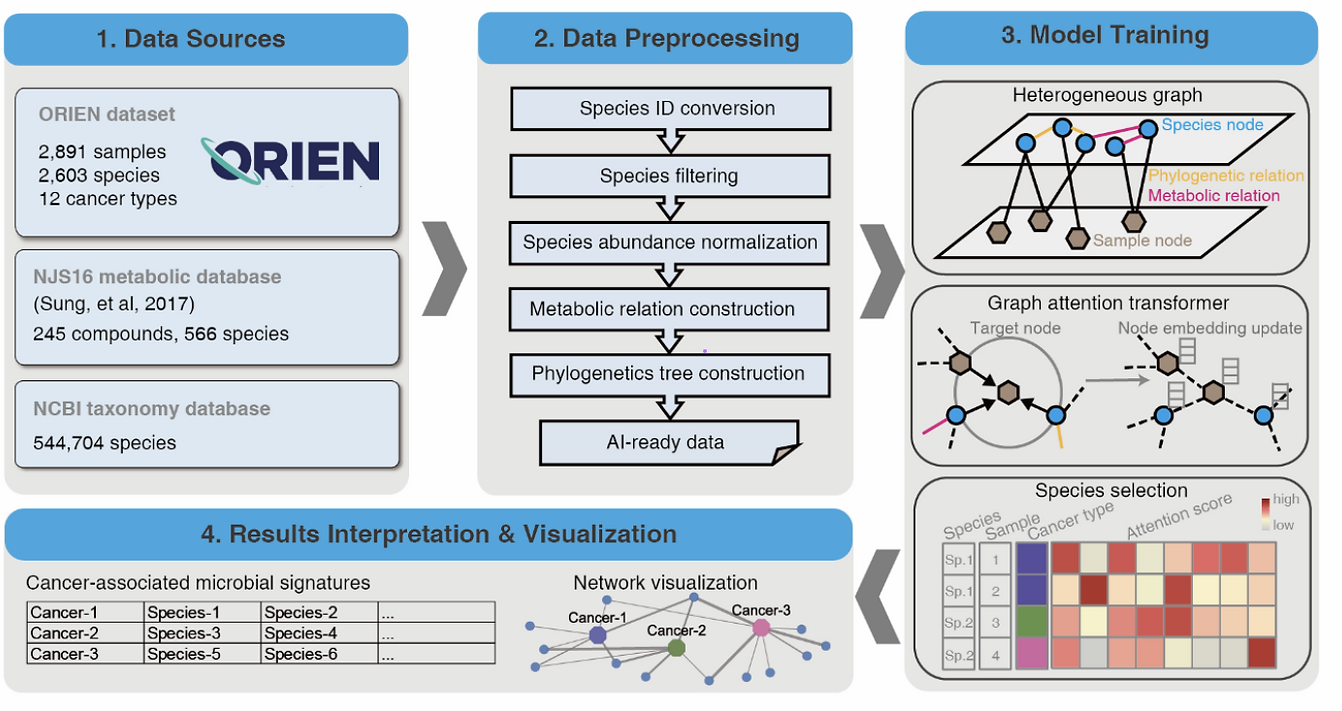
Evidence supports significant interactions among microbes, immune cells, and tumor cells in at least 10–20% of human cancers, emphasizing the importance of further investigating these complex relationships. However, the implications and significance of tumor-related microbes remain largely unknown. Studies have demonstrated the critical roles of host microbes in cancer prevention and treatment responses. Understanding interactions between host microbes and cancer can drive cancer diagnosis and microbial therapeutics (bugs as drugs). Computational identification of cancer-specific microbes and their associations is still challenging due to the high dimensionality and high sparsity of intratumoral microbiome data, which requires large datasets containing sufficient event observations to identify relationships, and the interactions within microbial communities, the heterogeneity in microbial composition, and other confounding effects that can lead to spurious associations. To solve these issues, we present a bioinformatics tool, MEGA, to identify the microbes most strongly associated with 12 cancer types. We demonstrate its utility on a dataset from a consortium of 9 cancer centers in the Oncology Research Information Exchange Network (ORIEN). This package has 3 unique features: species-sample relations are represented in a heterogeneous graph and learned by a graph attention network; it incorporates metabolic and phylogenetic information to reflect intricate relationships within microbial communities; and it provides multiple functionalities for association interpretations and visualizations. We analyzed 2704 tumor RNA-seq samples and MEGA interpreted the tissue-resident microbial signatures of each of 12 cancer types. MEGA can effectively identify cancer-associated microbial signatures and refine their interactions with tumors.



bIoRxiv
Caroline Wheeler, Samuel Coleman, Rebecca Hoyd, Louis Denko, Carlos Chan, Michelle Churchman, Nicholas Denko, Rebecca Dodd, Islam Eljilany, Sheetal Hardikar, Marium Husain, Alexandra Ikeguchi, Ning Jin, Qin Ma, Martin McCarter, Afaf Osman, Lary Robinson, Eric Singer, Gabriel Tinoco, Cornelia Ulrich, Yousef Zakharia, Daniel Spakowicz, Ahmad Tarhini, and Aik Choon Tan
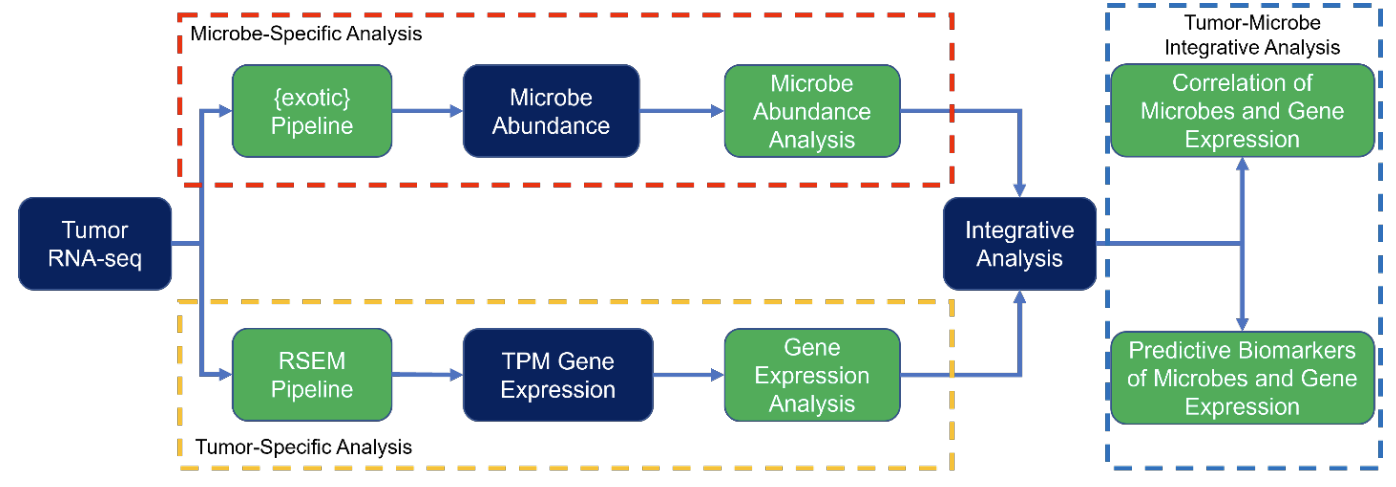
Emerging evidence supports the important role of the tumor microbiome in oncogenesis, cancer immune phenotype, cancer progression, and treatment outcomes in many malignancies. In this study, we investigated the metastatic melanoma tumor microbiome and potential roles in association with clinical outcomes, such as survival, in patients with metastatic disease treated with immune checkpoint inhibitors (ICIs). Baseline tumor samples were collected from 71 patients with metastatic melanoma before treatment with ICIs. Bulk RNA-seq was conducted on the formalin-fixed paraffin-embedded (FFPE) tumor samples. Durable clinical benefit (primary clinical endpoint) following ICIs was defined as overall survival ≥24 months and no change to the primary drug regimen (responders). We processed RNA-seq reads to carefully identify exogenous sequences using the {exotic}tool. The 71 patients with metastatic melanoma ranged in age from 24 to 83 years, 59% were male, and 55% survived >24 months following the initiation of ICI treatment. Exogenous taxa were identified in the tumor RNA-seq, including bacteria, fungi, and viruses. We found differences in gene expression and microbe abundances in immunotherapy responsive versus non-responsive tumors. Responders showed significant enrichment of several microbes including Fusobacterium nucleatum, and non-responders showed enrichment of fungi, as well as several bacteria. These microbes correlated with immune-related gene expression signatures. Finally, we found that models for predicting prolonged survival with immunotherapy using both microbe abundances and gene expression outperformed models using either dataset alone. Our findings warrant further investigation and potentially support therapeutic strategies to modify the tumor microbiome in order to improve treatment outcomes with ICIs.

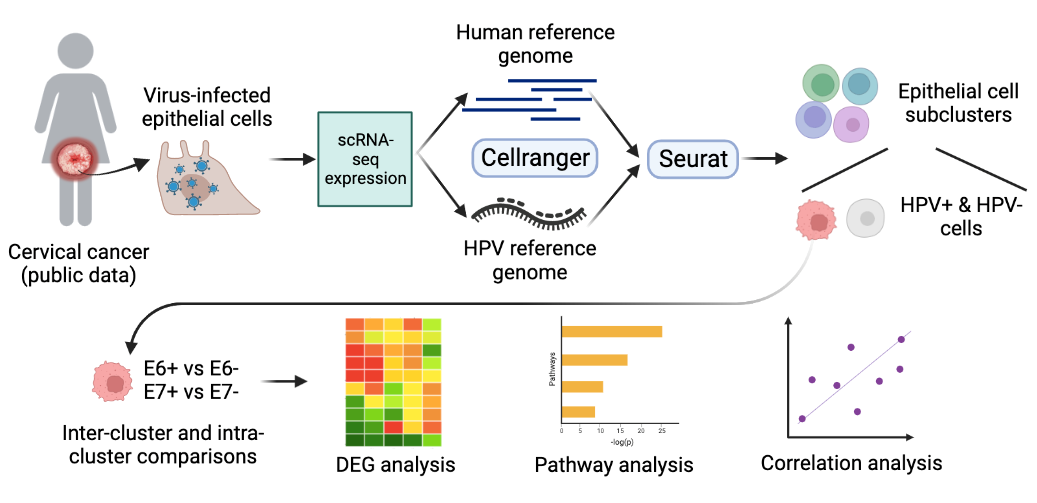
Identification of epithelial layer-specific markers from host single-cell transcriptomic in HPV-infected cervical cancer
In progress
Yingjie Li, Cankun Wang, Anjun Ma, Xuefeng Liu, Qin Ma
Tumor microbe associations with clinical outcomes in soft tissue sarcoma patients
In progress
Marium Husain, Ahmed Hussein, YunZhou Liu, Caroline E. Wheeler, Rebecca Hoyd, Lary A. Robinson, Carlos H.F. Chan, Yousef Zakharia, Rebecca D. Dodd, Cornelia M. Ulrich, Sheetal Hardikar, Michelle L. Churchman, Ahmad A. Tarhini, Eric A. Singer, Alexandra P. Ikeguchi, Martin D. McCarter, Nicholas Denko, Ning Jin, Afaf E.G. Osman, Aik Choon Tan, Qin Ma, Gregory Riedlinger, Bryan P. Schneider, Daniel Spakowicz, and Gabriel Tinoco
Metagenomics & metatranscriptomics
Reviews
Methods
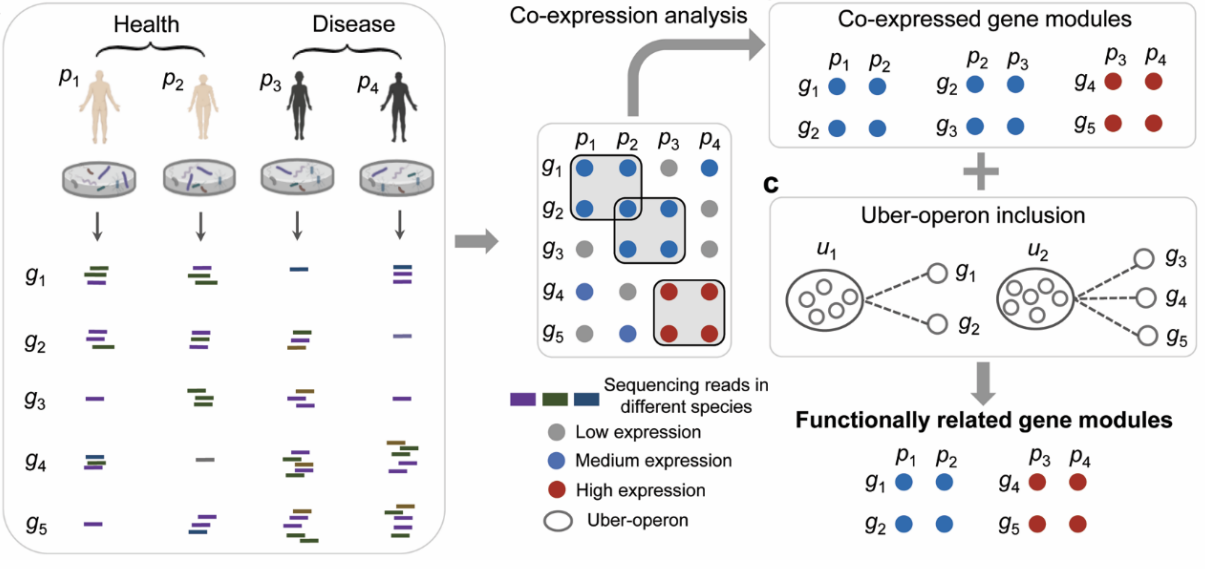
Microbial systems biology
Methods
Bacterial functional genomics
